MitoDNA™ Red 710
Product key features
- mtDNA-Specific: Selectively binds to mtDNA without nuclear interference
- Live-Cell Compatibility: Easily permeates cells for real-time mtDNA tracking
- High Signal Clarity: Large Stokes shift ensures a strong signal-to-noise ratio for multiplex imaging
- Broad Research Applicability: Ideal for investigating mtDNA-linked diseases and mitochondrial function
Product description
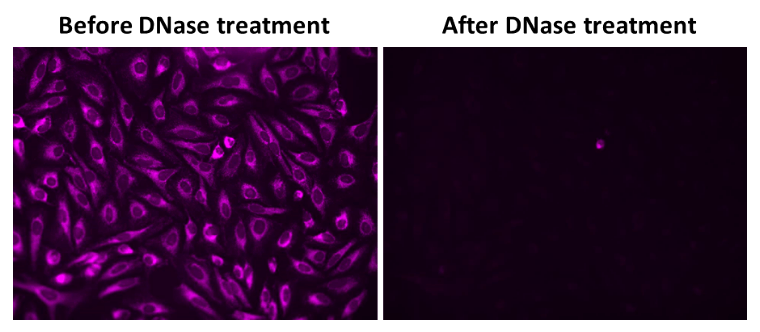
There are limited probes available that effectively detect mitochondrial DNA (mtDNA). Conventional fluorescent DNA probes, such as DAPI, Hoechst, or SYBR® Green, lack the specificity required for mitochondrial targeting and primarily stain nuclear DNA. MitoDNA™ Red 710 is a cell-permeable dye that selectively stains mtDNA in live cells, providing a method for dynamic imaging of mtDNA. This dye exhibits a large Stokes shift (Ex/Em = 510/710 nm), offering a high signal-to-noise ratio and enabling multiplex staining with other fluorescent probes. mtDNA is a small, circular DNA molecule located within the mitochondria in the cytoplasm. It supplements nuclear DNA and encodes 37 genes essential for mitochondrial and cellular functions. Mitochondria are responsible for ATP synthesis through oxidative phosphorylation and house the genetic information necessary for synthesizing key enzymes, transfer RNA (tRNA), and ribosomal RNA (rRNA). Mutations and disorders in mtDNA are implicated in a range of pathologies, including age-related hearing loss, diabetes, and organ dysfunctions in the brain, heart, and liver. Additionally, mtDNA mutations are associated with an elevated risk of various cancers, including lymphomas, leukemias, and tumors in the breast, intestines, liver, and kidneys.
Example protocol
AT A GLANCE
Before using MitoDNA™ Red 710 for the first time, allow it to thaw at room temperature. Then, briefly centrifuge it to collect the dried pellet.
Prepare cells in a growth medium.
Stain cells with MitoDNA™ Red 710 working solution.
Incubate samples for 5 to 15 minutes at 37 °C.
Monitor fluorescence intensity at Ex/Em = 510/710 nm.
PREPARATION OF STOCK SOLUTIONS
Unless otherwise noted, all unused stock solutions should be divided into single-use aliquots and stored at -20 °C after preparation. Avoid repeated freeze-thaw cycles
Prepare a 5 to 10 mM MitoDNA™ Red 710 stock solution in DMSO. For example, add 290 μL of DMSO to the MitoDNA™ Red 710 vial to create a 10 mM stock solution.
Note: Prepare a single aliquot of the unused MitoDNA™ Red 710 stock solution and store it at ≤ -20 º C, protected from light. Avoid repeated freeze-thaw cycles.
PREPARATION OF WORKING SOLUTION
Prepare a 5 to 10 μM working solution by diluting the MitoDNA™ Red 710 stock solution in Hanks' solution with 20 mM HEPES buffer (HHBS).
Note: For optimal results, use this solution within a few hours of preparation.
Note: Cover the working solution with foil or store it in a dark place to protect it from light.
SAMPLE EXPERIMENTAL PROTOCOL
Plate the cells in a 96-well plate with black walls and a clear bottom.
Remove the cell culture medium and add 100 µL of MitoDNA™ Red 710 working solution directly to the cells.
Incubate the cells at 37°C for 5-15 minutes, protected from light.
Note: The concentration and incubation time of MitoDNA™ Red 710 may vary depending on the cell line. Test different concentrations to determine the optimal dose.
Remove the dye working solution and wash the cells twice with HHBS buffer.
Add HHBS buffer and analyze the cells using a fluorescence microscope with excitation/emission settings of 510/710 nm.
Spectrum
Product family
| Name | Excitation (nm) | Emission (nm) |
| MitoDNA™ Red 610 | 508 | 607 |
| MitoDNA™ Red 680 | 597 | 681 |
References
Authors: Li, Wenting and Li, Yuting and Zhao, Jie and Liao, Jiabao and Wen, Weibo and Chen, Yao and Cui, Huantian
Journal: Pathology, research and practice (2024): 155330
Authors: Huang, Penghui and Li, Li and Chen, Yaohua and Li, Yuping and Zhu, Dan and Cui, Jian
Journal: Open life sciences (2024): 20220872
Authors: Bulduk, Bengisu K and Tortajada, Juan and Valiente-Pallejà, Alba and Callado, Luís F and Torrell, Helena and Vilella, Elisabet and Meana, J Javier and Muntané, Gerard and Martorell, Lourdes
Journal: Psychiatry research (2024): 115928
Authors: Weissensteiner, Hansi and Forer, Lukas and Kronenberg, Florian and Schönherr, Sebastian
Journal: Nucleic acids research (2024)
Authors: Feng, Yue and You, Yingqian and Li, Mengying and Guan, Xin and Fu, Ming and Wang, Chenming and Xiao, Yang and He, Meian and Guo, Huan
Journal: The Science of the total environment (2024): 173767